Milestone 5 - Project Results
Expected Results
This is an example for the expected results:
Diffusion Coefficient Plot
Plot of the diffusion coefficient:
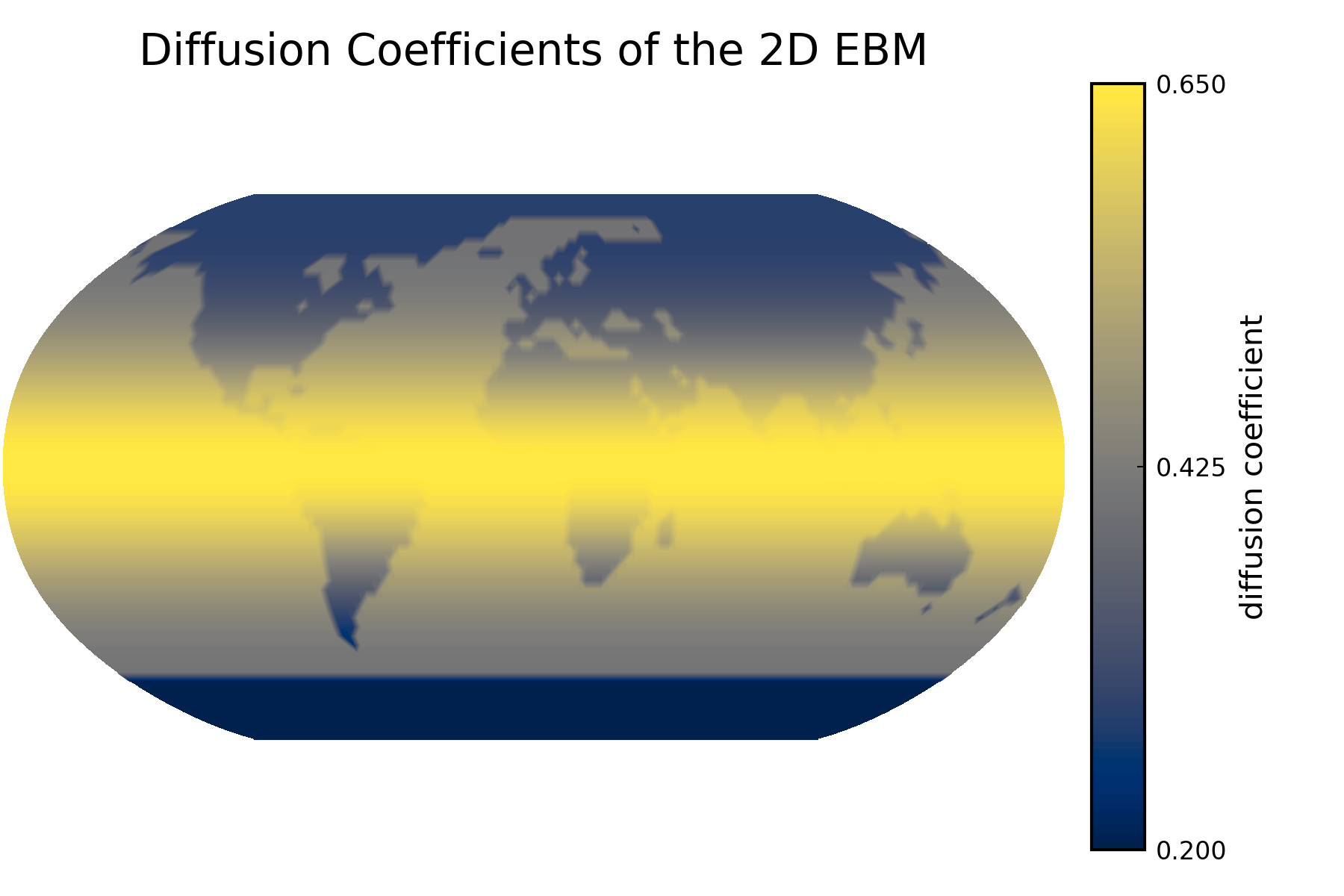

Sparsity Pattern of the Jacobian
Plot of the upper left corner of the sparsity pattern of the Jacobian. Only a the upper left part is plotted to make the pattern visible.
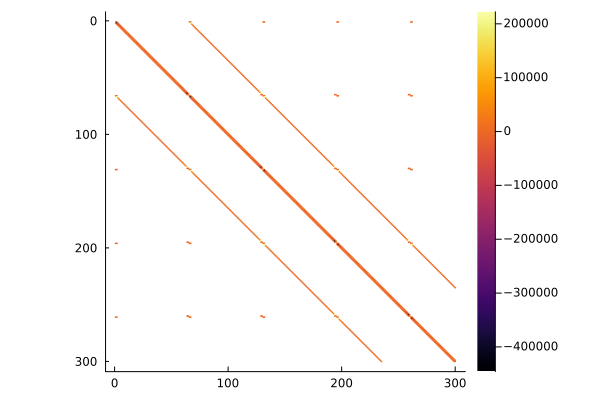
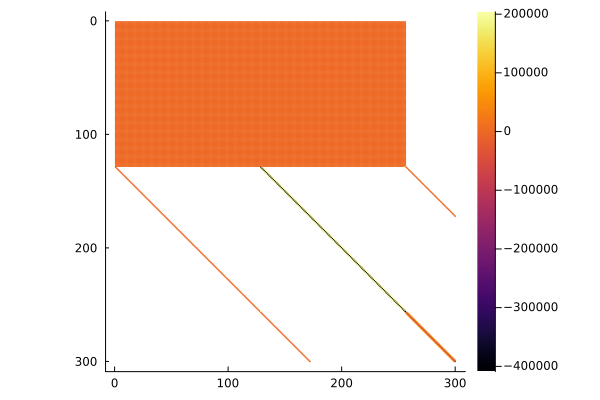
In a row-major language like Python, the sparsity pattern looks different:
⚠ Warning!
We strongly suggest to first work on the solutions on your own/within your group without checking directly for the reference solutions.
Files Download
The Python implementation of milestone 5 can be downloaded here (Julia version will be added soon):
Bonus - 3D plot
3D plot generated by Chong-Son Dröge, Fabian Höck, and Luca Levent Sommer.
Scripts for Milestone 5
You can also check out our Julia and Python implementations of milestone 5 in this site.
Julia implementation of milestone 5
include("milestone1.jl")
include("milestone2.jl")
include("milestone3.jl")
using Arpack: eigs
struct Mesh
n_latitude::Int
n_longitude::Int
ndof::Int
h::Float64
area::Vector{Float64}
geom::Float64
csc2::Vector{Float64}
cot::Vector{Float64}
function Mesh(geo_dat)
n_latitude, n_longitude = size(geo_dat)
ndof = n_latitude * n_longitude
h = pi / (n_latitude - 1)
area = calc_area(geo_dat)
geom = sin(0.5 * h) / area[1]
csc2 = [1 / sin(h * (j - 1))^2 for j in 2:(n_latitude - 1)]
cot = [1 / tan(h * (j - 1)) for j in 2:(n_latitude - 1)]
return new(n_latitude, n_longitude, ndof, h, area, geom, csc2, cot)
end
end
function calc_diffusion_coefficients(geo_dat)
nlatitude, nlongitude = size(geo_dat)
coeff_ocean_poles = 0.4
coeff_ocean_equator = 0.65
coeff_equator = 0.65
coeff_north_pole = 0.28
coeff_south_pole = 0.2
function diffusion_coefficient(j, i)
# Compute the j value of the equator
j_equator = Int((nlatitude - 1) / 2) + 1
theta = pi * (j - 1) / (nlatitude - 1)
colat = sin(theta)^5
geo = geo_dat[j, i]
if geo == 5 # ocean
return coeff_ocean_poles + (coeff_ocean_equator - coeff_ocean_poles) * colat
else # land, sea ice, etc
if j <= j_equator # northern hemisphere
# on the equator colat=1 -> coefficients for norhern/southern hemisphere cancels out
return coeff_north_pole + (coeff_equator - coeff_north_pole) * colat
else # southern hemisphere
return coeff_south_pole + (coeff_equator - coeff_south_pole) * colat
end
end
end
return [diffusion_coefficient(j, i) for j in 1:nlatitude, i in 1:nlongitude]
end
function calc_diffusion_operator(mesh, diffusion_coeff, temperature)
diffusion_op = similar(diffusion_coeff)
return calc_diffusion_operator!(diffusion_op, mesh, diffusion_coeff, temperature)
end
# Inplace version to avoid allocations
function calc_diffusion_operator!(diffusion_op, mesh, diffusion_coeff, temperature)
diffusion_op .= 0.0
# North Pole
factor = sin(mesh.h / 2) / (4 * pi * mesh.area[1])
# Use `@views` to avoid (most) allocations
@views diffusion_op[1, :] .= factor * 0.5 *
dot(diffusion_coeff[1, :] + diffusion_coeff[2, :],
temperature[2, :] - temperature[1, :])
# South Pole
factor = sin(mesh.h / 2) / (4 * pi * mesh.area[end])
# Use `@views` to avoid (most) allocations
@views diffusion_op[end, :] .= factor * 0.5 *
dot(diffusion_coeff[end, :] +
diffusion_coeff[end - 1, :],
temperature[end - 1, :] - temperature[end, :])
for i in 1:(mesh.n_longitude)
# Only loop over inner nodes
for j in 2:(mesh.n_latitude - 1)
# There are the special cases of i=1 and i=mesh.n_longitude.
# We have a periodic boundary condition, so for i=1, we want i-1 to be the last entry.
# For i=mesh.n_longitude, we want i+1 to be 1.
# For this, we define variables ip1 (i plus 1) abd im1 (i minus 1)
# to avoid duplicating code.
if i == mesh.n_longitude
ip1 = 1
else
ip1 = i + 1
end
if i == 1
im1 = mesh.n_longitude
else
im1 = i - 1
end
# Note that mesh.csc2 does not contain the values at the poles
factor1 = mesh.csc2[j - 1] / mesh.h^2
term1 = factor1 * (-2 * diffusion_coeff[j, i] * temperature[j, i] +
(diffusion_coeff[j, i] -
0.25 * (diffusion_coeff[j, ip1] - diffusion_coeff[j, im1])) *
temperature[j, im1] +
(diffusion_coeff[j, i] +
0.25 * (diffusion_coeff[j, ip1] - diffusion_coeff[j, im1])) *
temperature[j, ip1])
factor2 = 1 / mesh.h^2
term2 = factor2 * (-2 * diffusion_coeff[j, i] * temperature[j, i] +
(diffusion_coeff[j, i] -
0.25 *
(diffusion_coeff[j + 1, i] - diffusion_coeff[j - 1, i])) *
temperature[j - 1, i]
+
(diffusion_coeff[j, i] +
0.25 *
(diffusion_coeff[j + 1, i] - diffusion_coeff[j - 1, i])) *
temperature[j + 1, i])
term3 = mesh.cot[j - 1] * diffusion_coeff[j, i] * 0.5 / mesh.h *
(temperature[j + 1, i] - temperature[j - 1, i])
diffusion_op[j, i] = term1 + term2 + term3
end
end
return diffusion_op
end
function calc_operator_ebm_2d(temperature, mesh, diffusion_coeff, heat_capacity,
radiative_cooling_feedback=2.15)
diffusion_op = calc_diffusion_operator(mesh, diffusion_coeff, temperature)
return (diffusion_op .- radiative_cooling_feedback * temperature) ./ heat_capacity
end
# Inplace version to avoid allocations. Called from the Jacobian computation
function calc_operator_ebm_2d!(operator, temperature, mesh, diffusion_coeff, heat_capacity,
radiative_cooling_feedback=2.15)
calc_diffusion_operator!(operator, mesh, diffusion_coeff, temperature)
operator .-= radiative_cooling_feedback * temperature
operator ./= heat_capacity
return operator
end
function calc_source_terms_ebm_2d(heat_capacity, solar_forcing, radiative_cooling)
return (solar_forcing .- radiative_cooling) ./ heat_capacity
end
function calc_rhs_ebm_2d(temperature, mesh, diffusion_coeff, heat_capacity, solar_forcing,
radiative_cooling)
operator = calc_operator_ebm_2d(temperature, mesh, diffusion_coeff, heat_capacity)
source_terms = calc_source_terms_ebm_2d(heat_capacity, solar_forcing, radiative_cooling)
return operator + source_terms
end
function timestep_euler_forward_2d(temperature, t, delta_t,
mesh, diffusion_coeff, heat_capacity, solar_forcing,
radiative_cooling)
# Note that this function modifies the first argument instead of returning the result
if t == 1
t_old = size(temperature, 3)
else
t_old = t - 1
end
return temperature[:, :, t] = temperature[:, :, t_old] +
delta_t * calc_rhs_ebm_2d(temperature[:, :, t_old], mesh,
diffusion_coeff, heat_capacity,
solar_forcing[:, :, t_old],
radiative_cooling)
end
function compute_equilibrium_2d(timestep_function, mesh, diffusion_coeff, heat_capacity,
solar_forcing, radiative_cooling;
max_iterations=100, rel_error=2.0e-5, verbose=true,
initial_temperature=zeros((mesh.n_latitude,
mesh.n_longitude,
size(solar_forcing, 3))))
# Number of time steps per year
ntimesteps = size(solar_forcing, 3)
# Step size
delta_t = 1 / ntimesteps
temperature = initial_temperature
# Area-mean in every time step
area_mean_temp = zeros(ntimesteps)
# Average temperature over all time steps from the previous iteration to approximate the error
old_avg_temperature = 0
for i in 1:max_iterations
for t in 1:ntimesteps
timestep_function(temperature, t, delta_t,
mesh, diffusion_coeff, heat_capacity, solar_forcing,
radiative_cooling)
@views area_mean_temp[t] = calc_mean(temperature[:, :, t], mesh.area)
end
avg_temperature = sum(area_mean_temp) / ntimesteps
verbose && println("Average annual temperature in iteration $i is $avg_temperature")
if abs(avg_temperature - old_avg_temperature) < rel_error
# We can assume that the error is sufficiently small now.
verbose && println("Equilibrium reached!")
break
else
old_avg_temperature = avg_temperature
end
end
return temperature, area_mean_temp
end
function plot_diffusion_coefficient(diffusion_coeff)
vmin = minimum(diffusion_coeff)
vmax = maximum(diffusion_coeff)
nlatitude, nlongitude = size(diffusion_coeff)
x, y = robinson_projection(nlatitude, nlongitude)
p = contourf(x, y, diffusion_coeff,
clims=(vmin, vmax),
levels=LinRange(vmin, vmax, 100),
aspect_ratio=1,
title="Diffusion Coefficients of the 2D EBM",
c=:cividis,
colorbar_title="diffusion coefficient",
colorbar_ticks=([vmin, 0.5 * (vmin + vmax), vmax]),
axis=([], false),
dpi=300)
return p
end
function calc_jacobian_ebm_2d(mesh, diffusion_coeff, heat_capacity,
radiative_cooling_feedback=2.15)
jacobian = zeros((mesh.ndof, mesh.ndof))
test_temperature = zeros(size(diffusion_coeff))
op = similar(diffusion_coeff)
index = 1
for i in 1:(mesh.n_longitude), j in 1:(mesh.n_latitude)
test_temperature[j, i] = 1.0
# Use inplace version to avoid a lot of allocations
calc_operator_ebm_2d!(op, test_temperature, mesh, diffusion_coeff, heat_capacity,
radiative_cooling_feedback)
# Convert matrix to vector.
# Note that this must be compatible with the loop order, so that `index` is correct.
# `vec` works column-wise (as opposed to `flatten` in Python),
# so we must loop over columns first in order to get the correct Jacobian.
# To be precise, `vec(test_temperature)` must be the `index`-th unit vector.
jacobian[:, index] = vec(op)
# Reset test_temperature
test_temperature[j, i] = 0.0
index += 1
end
return jacobian
end
# Run code
function milestone5()
geo_dat = read_geography(joinpath(@__DIR__, "input", "The_World128x65.dat"))
mesh = Mesh(geo_dat)
albedo = calc_albedo(geo_dat)
heat_capacity = calc_heat_capacity(geo_dat)
# Compute solar forcing
true_longitude = read_true_longitude(joinpath(@__DIR__, "input", "True_Longitude.dat"))
solar_forcing = calc_solar_forcing(albedo, true_longitude)
# Compute and plot diffusion coefficient
diffusion_coeff = calc_diffusion_coefficients(geo_dat)
plot_diffusion_coeff = plot_diffusion_coefficient(diffusion_coeff)
co2_ppm = 315.0
radiative_cooling = calc_radiative_cooling_co2(co2_ppm)
compute_equilibrium_2d(timestep_euler_forward_2d, mesh, diffusion_coeff,
heat_capacity, solar_forcing,
radiative_cooling, max_iterations=2)
# The Jacobian has three diagonals of non-zero entries and two blocks of non-zero entries for the poles.
# We only show a subset of the entries, otherwise the two side diagonals are not visible.
jacobian = calc_jacobian_ebm_2d(mesh, diffusion_coeff, heat_capacity)
# Warning: Plotting the whole matrix with the PythonPlot backend is very slow.
# Use the GR backend when examining the whole Jacobian.
gr()
plot_jacobian = spy(jacobian[1:300, 1:300])
println("Done with Jacobian")
# This computes only the 6 eigenvalues with largest magnitude and no eigenvectors
eigenvalues, _ = eigs(jacobian, ritzvec=false)
# The maximum absolute value of the eigenvalues is too high for an efficient explicit time integration scheme.
print(max(maximum(real.(eigenvalues)), -minimum(real.(eigenvalues))))
# Show the plots
display(plot_diffusion_coeff)
display(plot_jacobian)
return plot_diffusion_coeff, plot_jacobian
endPython implementation of milestone 5
import matplotlib.pyplot as plt
import numpy as np
import scipy
from numba import njit
from milestone1 import read_geography, plot_robinson_projection
from milestone2 import calc_albedo, calc_heat_capacity, calc_solar_forcing, read_true_longitude
from milestone3 import calc_area, calc_mean, calc_radiative_cooling_co2
class Mesh:
def __init__(self, geo_dat):
self.n_latitude, self.n_longitude = geo_dat.shape
self.ndof = self.n_latitude * self.n_longitude
self.h = np.pi / (self.n_latitude - 1)
self.area = calc_area(geo_dat)
self.geom = np.sin(0.5 * self.h) / self.area[0]
self.csc2 = np.array([1 / np.sin(self.h * j) ** 2 for j in range(1, self.n_latitude - 1)])
self.cot = np.array([1 / np.tan(self.h * j) for j in range(1, self.n_latitude - 1)])
def calc_diffusion_coefficients(geo_dat):
nlatitude, nlongitude = geo_dat.shape
coeff_ocean_poles = 0.40
coeff_ocean_equator = 0.65
coeff_equator = 0.65
coeff_north_pole = 0.28
coeff_south_pole = 0.20
def diffusion_coefficient(j, i):
# Compute the j value of the equator
j_equator = int(nlatitude / 2)
theta = np.pi * j / np.real(nlatitude - 1)
colat = np.sin(theta) ** 5
geo = geo_dat[j, i]
if geo == 5: # ocean
return coeff_ocean_poles + (coeff_ocean_equator - coeff_ocean_poles) * colat
else: # land, sea ice, etc
if j <= j_equator: # northern hemisphere
# on the equator colat=1 -> coefficients for norhern/southern hemisphere cancels out
return coeff_north_pole + (coeff_equator - coeff_north_pole) * colat
else: # southern hemisphere
return coeff_south_pole + (coeff_equator - coeff_south_pole) * colat
return np.array(
[[diffusion_coefficient(j, i) for i in range(nlongitude)] for j in range(nlatitude)]
)
def calc_diffusion_operator(mesh, diffusion_coeff, temperature):
h = mesh.h
area = mesh.area
n_latitude = mesh.n_latitude
n_longitude = mesh.n_longitude
csc2 = mesh.csc2
cot = mesh.cot
return calc_diffusion_operator_inner(h, area, n_latitude, n_longitude, csc2, cot, diffusion_coeff, temperature)
# This function is slow. It takes about 22ms on my machine, which is a lot for such a simple function.
# The problem is that Python is an interpreted (as opposed to compiled) language and therefore very slow.
# We could try to outsource most of that code to Numpy functions to speed things up, but that is not easy.
# Instead, we will use Numba, which is a JIT (just-in-time) compiler for Python to compile this function to fast
# machine code.
# With Numba, we can just add `@njit` to the function to tell Numba to compile it directly to machine code.
# Note, however, that this does not work with classes like our `Mesh`.
# Therefore, we cannot pass `mesh` to this function. Instead, we have to unpack all fields of `Mesh` above and
# add a wrapper function that does the unpacking.
#
# Times on my machine:
# Without `@njit`: 22ms (185s total for 8320 calls when calculating the Jacobian).
# With `@njit`: 25µs (209ms total for 8320 calls when calculating the Jacobian).
#
# This gives us a ~1000x speedup! Just by slightly rewriting the function and adding a decorator.
# Of course, we could try to further optimize this function, but that would be a waste of time, as it's not
# performance-relevant anymore.
@njit
def calc_diffusion_operator_inner(h, area, n_latitude, n_longitude, csc2, cot, diffusion_coeff, temperature):
result = np.zeros(diffusion_coeff.shape)
# North Pole
factor = np.sin(h / 2) / (4 * np.pi * area[0])
result[0, :] = factor * 0.5 * np.dot(diffusion_coeff[0, :] + diffusion_coeff[1, :],
temperature[1, :] - temperature[0, :])
# South Pole
factor = np.sin(h / 2) / (4 * np.pi * area[-1])
result[-1, :] = factor * 0.5 * np.dot(diffusion_coeff[-1, :] + diffusion_coeff[-2, :],
temperature[-2, :] - temperature[-1, :])
for i in range(n_longitude):
# Only loop over inner nodes
for j in range(1, n_latitude - 1):
# There are the special cases of i=0 and i=n_longitude-1.
# We have a periodic boundary condition, so for i=0, we want i-1 to be the last entry.
# This happens automatically in Python when i=-1.
# For i=n_longitude-1, we want i+1 to be 0.
# For this, we define a variable ip1 (i plus 1) to avoid duplicating code.
if i == n_longitude - 1:
ip1 = 0
else:
ip1 = i + 1
# Note that csc2 does not contain the values at the poles
factor1 = csc2[j - 1] / h ** 2
term1 = factor1 * (-2 * diffusion_coeff[j, i] * temperature[j, i] +
(diffusion_coeff[j, i] - 0.25 * (diffusion_coeff[j, ip1] - diffusion_coeff[j, i - 1])) *
temperature[j, i - 1] +
(diffusion_coeff[j, i] + 0.25 * (diffusion_coeff[j, ip1] - diffusion_coeff[j, i - 1])) *
temperature[j, ip1])
factor2 = 1 / h ** 2
term2 = factor2 * (-2 * diffusion_coeff[j, i] * temperature[j, i] +
(diffusion_coeff[j, i] - 0.25 *
(diffusion_coeff[j + 1, i] - diffusion_coeff[j - 1, i])) *
temperature[j - 1, i]
+ (diffusion_coeff[j, i] + 0.25 *
(diffusion_coeff[j + 1, i] - diffusion_coeff[j - 1, i])) *
temperature[j + 1, i])
term3 = cot[j - 1] * diffusion_coeff[j, i] * 0.5 / h * (temperature[j + 1, i] - temperature[j - 1, i])
result[j, i] = term1 + term2 + term3
return result
def calc_operator_ebm_2d(temperature, mesh, diffusion_coeff, heat_capacity, radiative_cooling_feedback=2.15):
diffusion_op = calc_diffusion_operator(mesh, diffusion_coeff, temperature)
return (diffusion_op - radiative_cooling_feedback * temperature) / heat_capacity
def calc_source_terms_ebm_2d(heat_capacity, solar_forcing, radiative_cooling):
return (solar_forcing - radiative_cooling) / heat_capacity
def calc_rhs_ebm_2d(temperature, mesh, diffusion_coeff, heat_capacity, solar_forcing, radiative_cooling):
operator = calc_operator_ebm_2d(temperature, mesh, diffusion_coeff, heat_capacity)
source_terms = calc_source_terms_ebm_2d(heat_capacity, solar_forcing, radiative_cooling)
return operator + source_terms
def timestep_euler_forward_2d(temperature, t, delta_t,
mesh, diffusion_coeff, heat_capacity, solar_forcing, radiative_cooling):
# Note that this function modifies the first argument instead of returning the result
temperature[:, :, t] = temperature[:, :, t - 1] + delta_t * calc_rhs_ebm_2d(temperature[:, :, t - 1], mesh,
diffusion_coeff, heat_capacity,
solar_forcing[:, :, t - 1],
radiative_cooling)
def compute_equilibrium_2d(timestep_function, mesh, diffusion_coeff, heat_capacity, solar_forcing, radiative_cooling,
max_iterations=100, rel_error=2e-5, verbose=True, initial_temperature=None):
if initial_temperature is None:
# We start with a constant temperature of 0 in every grid point throughout the year
temperature = np.zeros((mesh.n_latitude, mesh.n_longitude, solar_forcing.shape[2]))
else:
temperature = initial_temperature
# Number of time steps per year
ntimesteps = solar_forcing.shape[2]
# Step size
delta_t = 1 / ntimesteps
# Area-mean in every time step
area_mean_temp = np.zeros(ntimesteps)
# Average temperature over all time steps from the previous iteration to approximate the error
old_avg_temperature = 0
for i in range(max_iterations):
for t in range(ntimesteps):
timestep_function(temperature, t, delta_t,
mesh, diffusion_coeff, heat_capacity, solar_forcing, radiative_cooling)
area_mean_temp[t] = calc_mean(temperature[:, :, t], mesh.area)
avg_temperature = np.sum(area_mean_temp) / ntimesteps
verbose and print(f"Average annual temperature in iteration {i} is {avg_temperature}")
if np.abs(avg_temperature - old_avg_temperature) < rel_error:
# We can assume that the error is sufficiently small now.
verbose and print("Equilibrium reached!")
break
else:
old_avg_temperature = avg_temperature
return temperature, area_mean_temp
def plot_diffusion_coefficient(diffusion_coeff):
vmin = np.amin(diffusion_coeff)
vmax = np.amax(diffusion_coeff)
levels = np.linspace(vmin, vmax, 100)
# Reuse plotting function from milestone 1.
cbar = plot_robinson_projection(diffusion_coeff, "Diffusion Coefficients of the 2D EBM",
levels=levels, cmap="cividis",
vmin=vmin, vmax=vmax)
cbar.set_ticks([vmin, 0.5 * (vmin + vmax), vmax])
cbar.set_label("diffusion coefficient")
# Adjust size of plot to viewport to prevent clipping of the legend.
plt.tight_layout()
plt.show()
def calc_jacobian_ebm_2d(mesh, diffusion_coeff, heat_capacity, radiative_cooling_feedback=2.15):
jacobian = np.zeros((mesh.ndof, mesh.ndof))
test_temperature = np.zeros(diffusion_coeff.shape)
index = 0
for j in range(mesh.n_latitude):
for i in range(mesh.n_longitude):
test_temperature[j, i] = 1.0
op = calc_operator_ebm_2d(test_temperature, mesh, diffusion_coeff, heat_capacity,
radiative_cooling_feedback=radiative_cooling_feedback)
# Convert matrix to vector.
# Note that this must be compatible with the loop order, so that `index` is correct.
# `flatten` works row-wise, so we must loop over rows first in order to get the correct Jacobian.
# To be precise, `test_temperature.flatten()` must be the `index`-th unit vector.
jacobian[:, index] = op.flatten()
# Reset test_temperature
test_temperature[j, i] = 0.0
index += 1
return jacobian
# Run code
if __name__ == '__main__':
geo_dat_ = read_geography("input/The_World128x65.dat")
mesh_ = Mesh(geo_dat_)
albedo = calc_albedo(geo_dat_)
heat_capacity_ = calc_heat_capacity(geo_dat_)
# Compute solar forcing
true_longitude = read_true_longitude("input/True_Longitude.dat")
solar_forcing_ = calc_solar_forcing(albedo, true_longitude)
# Compute and plot diffusion coefficient
diffusion_coeff_ = calc_diffusion_coefficients(geo_dat_)
plot_diffusion_coefficient(diffusion_coeff_)
co2_ppm = 315.0
radiative_cooling_ = calc_radiative_cooling_co2(co2_ppm)
compute_equilibrium_2d(timestep_euler_forward_2d, mesh_, diffusion_coeff_, heat_capacity_, solar_forcing_,
radiative_cooling_, max_iterations=2)
# The Jacobian has three diagonals of non-zero entries and two blocks of non-zero entries for the poles.
# We only show a subset of the entries, otherwise the two side diagonals are not visible.
jacobian_ = calc_jacobian_ebm_2d(mesh_, diffusion_coeff_, heat_capacity_)
plt.spy(jacobian_[0:300, 0:300])
plt.show()
# Use Scipy to efficiently compute the eigenvalues of this sparse matrix.
# Converting the matrix to a sparse format first makes `eigs` ~100x faster.
eigenvalues = scipy.sparse.linalg.eigs(scipy.sparse.csr_matrix(jacobian_), return_eigenvectors=False)
# The maximum absolute value of the eigenvalues is too high for an efficient explicit time integration scheme.
print(np.real(max(max(eigenvalues), -min(eigenvalues))))Created by Gregor Gassner and Andrés Rueda-Ramírez with contributions by Simone Chiocchetti, Daniel Bach, Sophia Horak, Philipp Baasch, Benjamin Bolm, Erik Faulhaber, and Luca Sommer. Last modified: April 02, 2026. Website built with Franklin.jl and the Julia programming language.
